Join us! Three open positions
We currently have three fully financed openings for two postdocs and one PhD student. You can find the descriptions of the positions here: PhD position on nanoplasmonics for ultrafast single-molecule...
Peter Zijlstra Group

DOI: doi.org/10.1039/C9NR10218C
Matěj Horáček, Dion J. Engels, Peter Zijlstra
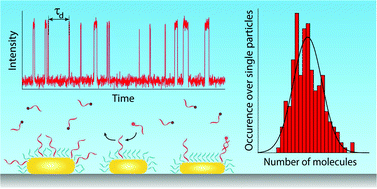
Applications of colloidal particles in the fields of i.e. biosensors, molecular targeting, or drug-delivery require their functionalization with biologically active and specific molecular ligands. Functionalization protocols often result in a heterogeneous population of particles with a varying density, spatial distribution and orientation of the functional groups on the particle surface. A lack of methods to directly resolve these molecular properties of the particle’s surface hampers optimization of functionalization protocols and applications. Here quantitative single-molecule interaction kinetics is used to count the number of ligands on the surface of hundreds of individual nanoparticles simultaneously. By analyzing the waiting-time between single-molecule binding events we quantify the particle functionalization both accurately and precisely for a large range of ligand densities. We observe significant particle-to-particle differences in functionalization which are dominated by the particle-size distribution for high molecular densities, but are substantially broadened for sparsely functionalized particles. From time-dependent studies we find that ligand reorganization on long timescales drastically reduces this heterogeneity, a process that has remained hidden up to now in ensemble-averaged studies. The quantitative single-molecule counting therefore provides a direct route to quantification and optimization of coupling protocols towards molecularly controlled colloidal interfaces.