Plasmon-Enhanced Single-Molecule Enzymology
Yuyang Wang and Peter Zijlstra
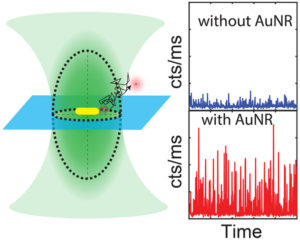
We present a numerical study on plasmon-enhanced single-molecule enzymology. We combine Brownian dynamics and electromagnetic simulations to calculate the enhancement of fluorescence signals of fluorogenic substrate converted by an enzyme conjugated to a plasmonic particle. We simulate the Brownian motion of a fluorescent product away from the active site of the enzyme, and calculate the photon detection rate taking into account modifications of the excitation and emission processes by coupling to the plasmon. We show that plasmon enhancement can boost the signal-to-noise ratio (SNR) of single turnovers by up to 100 fold compared to confocal microscopy. This enhancement factor is a trade-off between the reduced residence time in the near-field of the particle, and the enhanced emission intensity due to coupling to the plasmon. The enhancement depends on the size, shape and material of the particle and the photophysical properties of the fluorescent product. Our study provides guidelines on how to enhance the SNR of single-molecule enzyme studies and may aid in further understanding and quantifying static and dynamic heterogeneity.
Related Articles
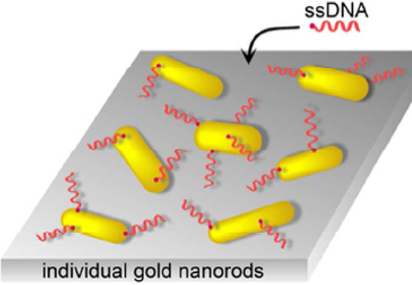
Heterogeneous kinetics in the functionalization of single plasmonic nanoparticles Matej Horacek, Rachel E. Armstrong, and Peter Zijlstra DOI: 10.1021/acs.langmuir.7b04027 The functionalization of gold nanoparticles with DNA has been studied extensively in solution, however...
We are excited to announce that a PhD vacancy is available for application at the molecular plasmonics group in Eindhoven University of Technology. PhD position on single-molecule plasmon sensing The...
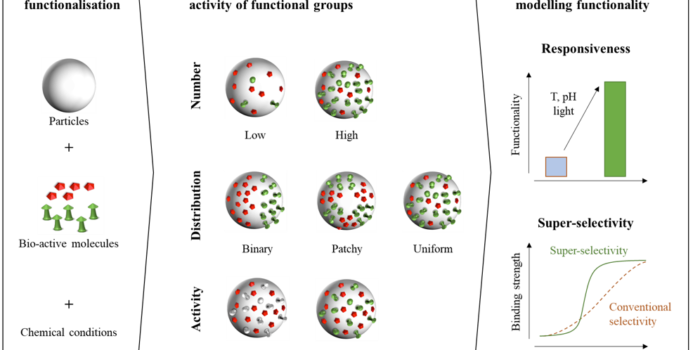
We are happy that our ITN grant called SuperCol was awarded! This is an international training network funded by the European Union and is coördinated by Peter. The network will...