Plasmon-Enhanced Single-Molecule Enzymology
Yuyang Wang and Peter Zijlstra
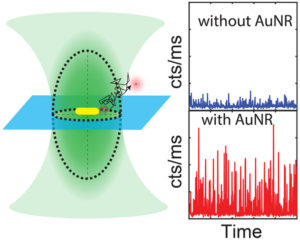
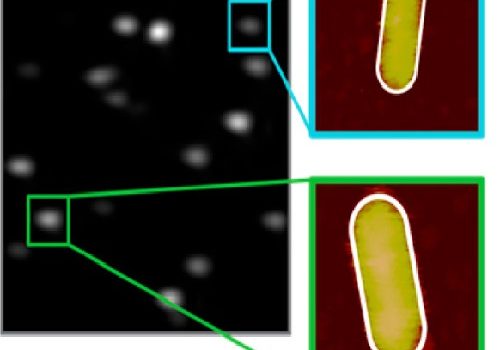
We present a numerical study on plasmon-enhanced single-molecule enzymology. We combine Brownian dynamics and electromagnetic simulations to calculate the enhancement of fluorescence signals of fluorogenic substrate converted by an enzyme conjugated to a plasmonic particle. We simulate the Brownian motion of a fluorescent product away from the active site of the enzyme, and calculate the photon detection rate taking into account modifications of the excitation and emission processes by coupling to the plasmon. We show that plasmon enhancement can boost the signal-to-noise ratio (SNR) of single turnovers by up to 100 fold compared to confocal microscopy. This enhancement factor is a trade-off between the reduced residence time in the near-field of the particle, and the enhanced emission intensity due to coupling to the plasmon. The enhancement depends on the size, shape and material of the particle and the photophysical properties of the fluorescent product. Our study provides guidelines on how to enhance the SNR of single-molecule enzyme studies and may aid in further understanding and quantifying static and dynamic heterogeneity.
Related Articles
On November 8th-11th,2021 the Max Planck Institute for Polymer Research (MPIP) hosted the workshop ‘Colloidal synthesis and colloid characterization’ which kicked off the next phase of SUPERCOL. Read a full...
The ITN SuperCol (a European consortium coordinated by Peter Zijlstra) contributes to an exhibition themed ‘Sensing the Invisible’ in a joint exhibit with students and researchers of the ITN Consense,...
Spatially Resolved Sensitivity of Single-Particle Plasmon Sensors Michael A. Beuwer, Bas van Hoof, and Peter Zijlstra DOI:10.1021/acs.jpcc.8b00849 The high sensitivity of localized surface plasmon resonance sensors to the local refractive...